 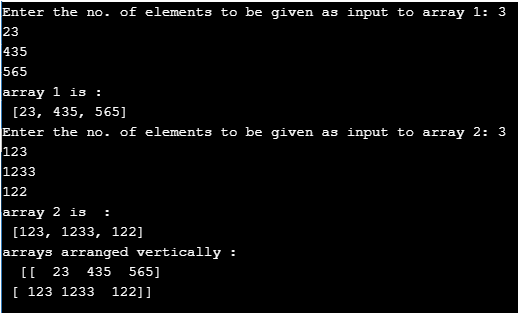
Np.dstack((gray,gray,gray)) will have shape (n0, n1,3). Np.hstack((gray,gray,gray)) will have shape (n0, n1*3), you can also do it by np.concatenate((gray,gray,gray),axis=1) Np.vstack((gray,gray,gray)) will have shape (n0*3, n1), you can also do it by np.concatenate((gray,gray,gray),axis=0) Simply put, numpy's functions are much more powerful. 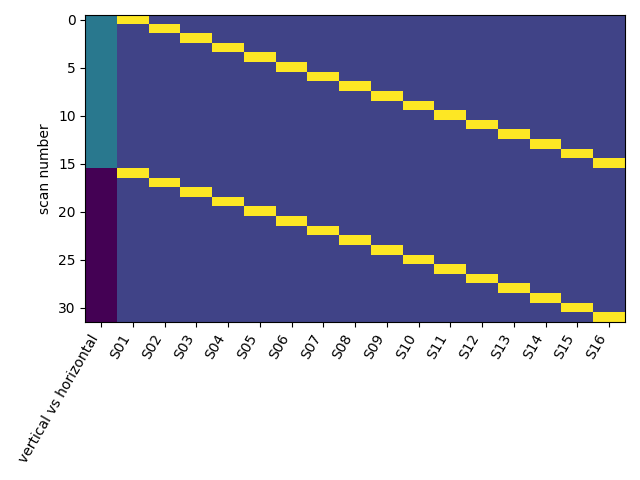
Obviously, To add multiple elements, you will use extend. Each time you add an element to the list. example see:Īppend is a function for python's built-in data structure list. Numpy.hstack: Stack arrays in sequence horizontally (column wise).Equivalent to np.concatenate(tup, axis=1), except for 1-D arrays where it concatenates along the first axis. Numpy.vstack: stack arrays in sequence vertically (row wise).Equivalent to np.concatenate(tup, axis=0) example see: My article Numpy Illustrated for details. If you're stacking two vectors, you've got three options:Īnd concatenate in its raw form is useful for 3D and above, see If you're stacking a matrice and a vector, hstack becomes tricky to use, so column_stack is a better option: If you have two matrices, you're good to go with just hstack and vstack: It, in effect, recursively concatenates along the nested lists. With axis=0, the effect is the same as np.array.īlock provide more general stacking and concatenation operations. That is, it expands the dims of all inputs (a bit like np.expand_dims), and then concatenates. Sl = (slice(None),) * axis + (_nx.newaxis,)Įxpanded_arrays = for arr in arrays]Ĭoncatenate(expanded_arrays, axis=axis, out=out) Shapes = set(arr.shape for arr in arrays)Īxis = normalize_axis_index(axis, result_ndim) They are just convenience functions.Īnd newer np.stack: arrays = In other words, they all work by tweaking the dimensions of the input arrays, and then concatenating on the right axis. turn all inputs in to 2d (or more) and concatenate on firstĬoncatenate(, axis=)Īrray(arr, copy=False, subok=True, ndmin=2).TĬoncatenate((asarray(arr), values), axis=axis) If not, here's a summary of their code: vstackĬoncatenate(, 0) The error your are getting, is anycodings_numpy-ndarray telling you precisely this.All the functions are written in Python except np.concatenate. 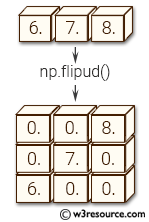
Now you can do both hstack() and anycodings_numpy-ndarray vstack() because a and b do have the anycodings_numpy-ndarray same shape, but what is the condition on anycodings_numpy-ndarray the shapes if they are not the same?įor vstack, the second dimension (index anycodings_numpy-ndarray 1) must match, while for hstack, it is anycodings_numpy-ndarray the first dimension (index 0) that must anycodings_numpy-ndarray match. Plt.ylabel('petal length ')Īssume we have two arrays of shape (2, anycodings_numpy-ndarray 3) each, say: a = np.array(, ])ī = np.array(, ])īoth hstack() and vstack() would stack anycodings_numpy-ndarray the two arrays, but along different anycodings_numpy-ndarray dimensions: np.vstack((a, b)) you can grow them using hstack and vstack although I am not sure how. Plot_decision_regions(X=X_combined_std, y=y_combined, classifier=ppn, test_idx=range(105,150)) One big win here over numpy is that theres the push function on Julia. X_combined_std= np.vstack((X_train_std, X_test_std)) Plt.scatter(X_test, X_test, c='', edgecolor= 'black', alpha= 0.9, linewidth=1, marker='o', s=100, label='test set' ) Plt.scatter (x=X, y= X, alpha=0.8, c=colors, marker= markers, label = cl, edgecolor = 'black') Plt.contourf(xx1, xx2, Z, alpha= 0.3, cmap = cmap) Xx1, xx2= np.meshgrid (np.arange(x1_min, x1_max, resolution), np.arange(x2_min, x2_max, resolution)) Ppn= Perceptron( max_iter=40,eta0= 0.1, random_state=1)ĭef plot_decision_regions(X, y, classifier,test_idx=None, resolution = 0.02):Ĭolors = ('red', 'blue', 'lightgreen', 'gray', 'cyan')Ĭmap = ListedColormap(colors) X_train, X_test, y_train, y_test= train_test_split(X, y, test_size=0.3, random_state=1, stratify=y) Match exactly, but along dimension 1, the array at index 0 has size 105īut I don't get that error when I use anycodings_numpy-ndarray np.hstack, why this happens? iris = datasets.load_iris() I get an error when I use anycodings_numpy-ndarray np.vstack ValueError:all the input array dimensions for the concatenation axis must I don't understand why I should use anycodings_numpy-ndarray np.hstack to adjust vector y y_combined=np.hstack((y_train, y_test)) 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
